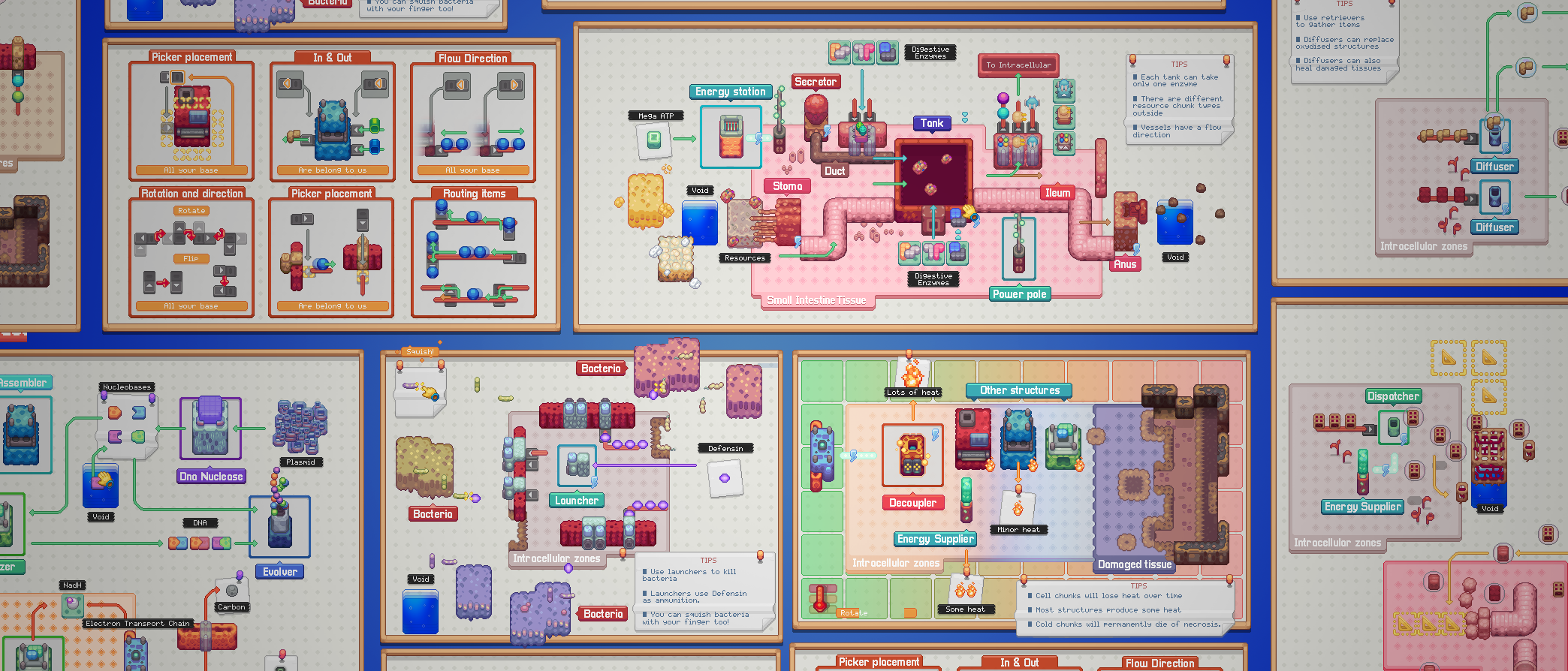
Circulation, Scale & UI improvements
Pathways documentation, extracellular circulation, game scale experiments and interactive tooltip work.



Start in a primordial broth as a single cell and make your way into life by creating the production chains needed to build, expand and evolve your initial organism.
 Survival, sandbox, automation
Survival, sandbox, automation
Lifecraft is about surviving: you begin in a primordial broth as a single cell and create the production chains needed to make your way into life.
The long vision is to evolve from a simple cell to a creature with organs: circulatory, respiratory, digestive and nervous systems, built by expanding and evolving the organism.
 Genre
Simulation / Strategy
Genre
Simulation / Strategy
 Mode
Single-player
Mode
Single-player
 Steam
33 achievements
Steam
33 achievements
 Status
Early Access
Status
Early Access
 Production
Ribosomes / Assemblers
Production
Ribosomes / Assemblers
 Routing
Microtubules / Pickers
Routing
Microtubules / Pickers
 Energy
ATP / ADP loop
Energy
ATP / ADP loop
 Threats
Temperature / Bacteria
Threats
Temperature / Bacteria
The in-game guide frames Lifecraft around production, routing, resources, energy, evolution, expansion, temperature, bacteria and specialized tissues.

Energy is generated by burning ATP into ADP; suppliers burn ATP and distribute energy to nearby buildings.
Open section
Evolvers consume nucleobases to discover new technologies, from production buildings to organelles.
Open section
Expansion creates space for buildings and scavenging resources, using dispatchers and the chunk planner.
Open section
Route items with microtubules and pickers, then use vessels and capillaries for extracellular flow.
Open section
Bacteria live in the broth, grow in colonies and can be answered with launchers, defensins and hemocytes.
Open section
Temperature, infection and nearby necrosis can reduce chunk health, so survival means preventing collapse.
Open section
Diffusers can deliver materials locally, replace oxidized buildings and heal infected tiles by supplying tissues.
Open section
Oxygen accepts electrons inside mitochondria; carbon dioxide is a waste product of energy reactions.
Open section
Stomas, tanks, ducts, enzymes and ileum vessels turn complex resources into absorbable nutrients.
Open section
Disposers throw unwanted items outside the cell; lysosomes dismantle items into raw materials.
Open section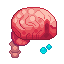
The pathways view shows interactions between structures, helping you read and automate the organism.
Open section
Assemblers, pickers, microtubules and logic structures turn repeated reactions into stable production chains.
Open section Screenshots
Screenshots
Browse cellular factories, routing lines, evolution screens, energy networks and the structures that make the organism work.




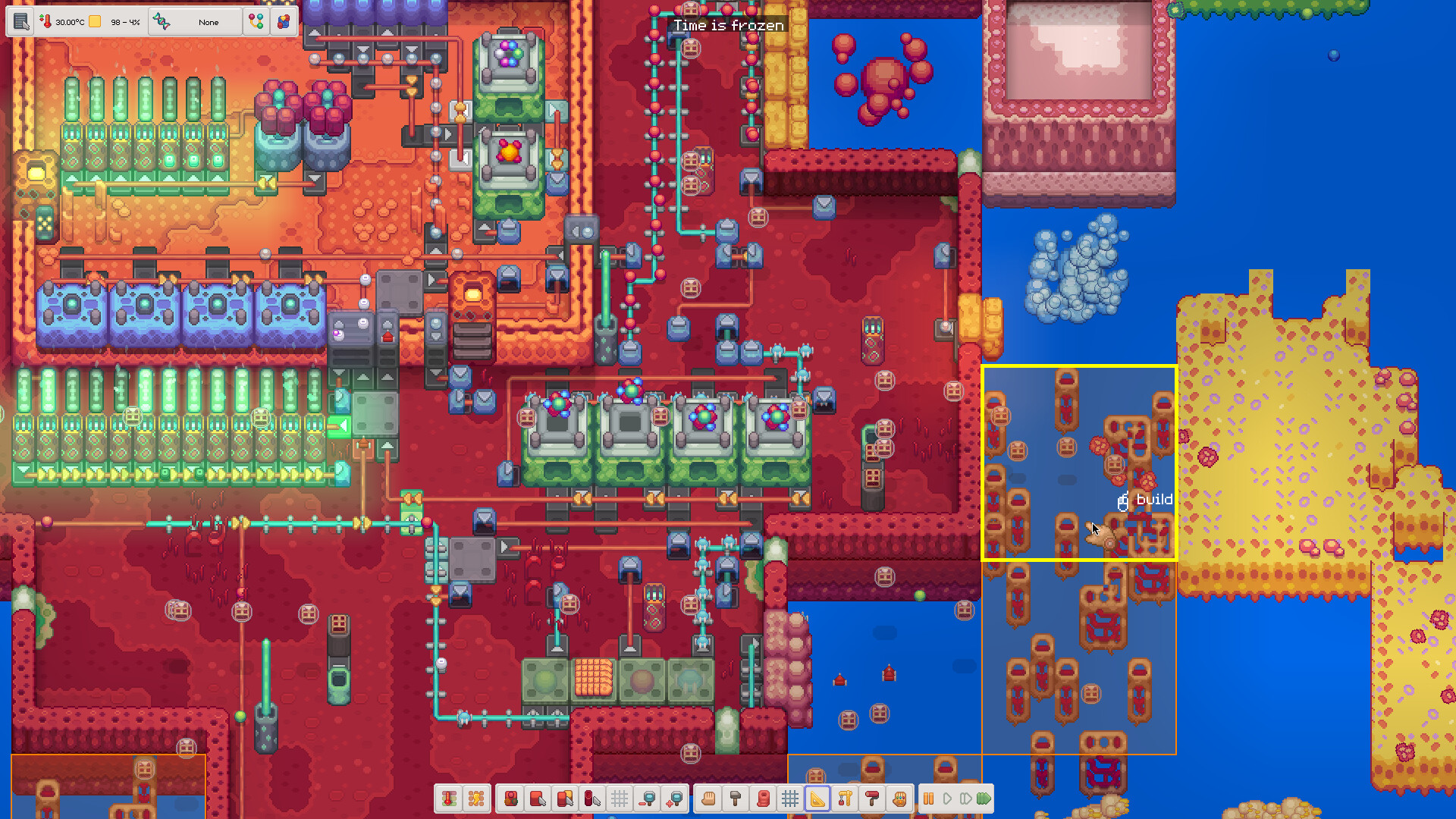
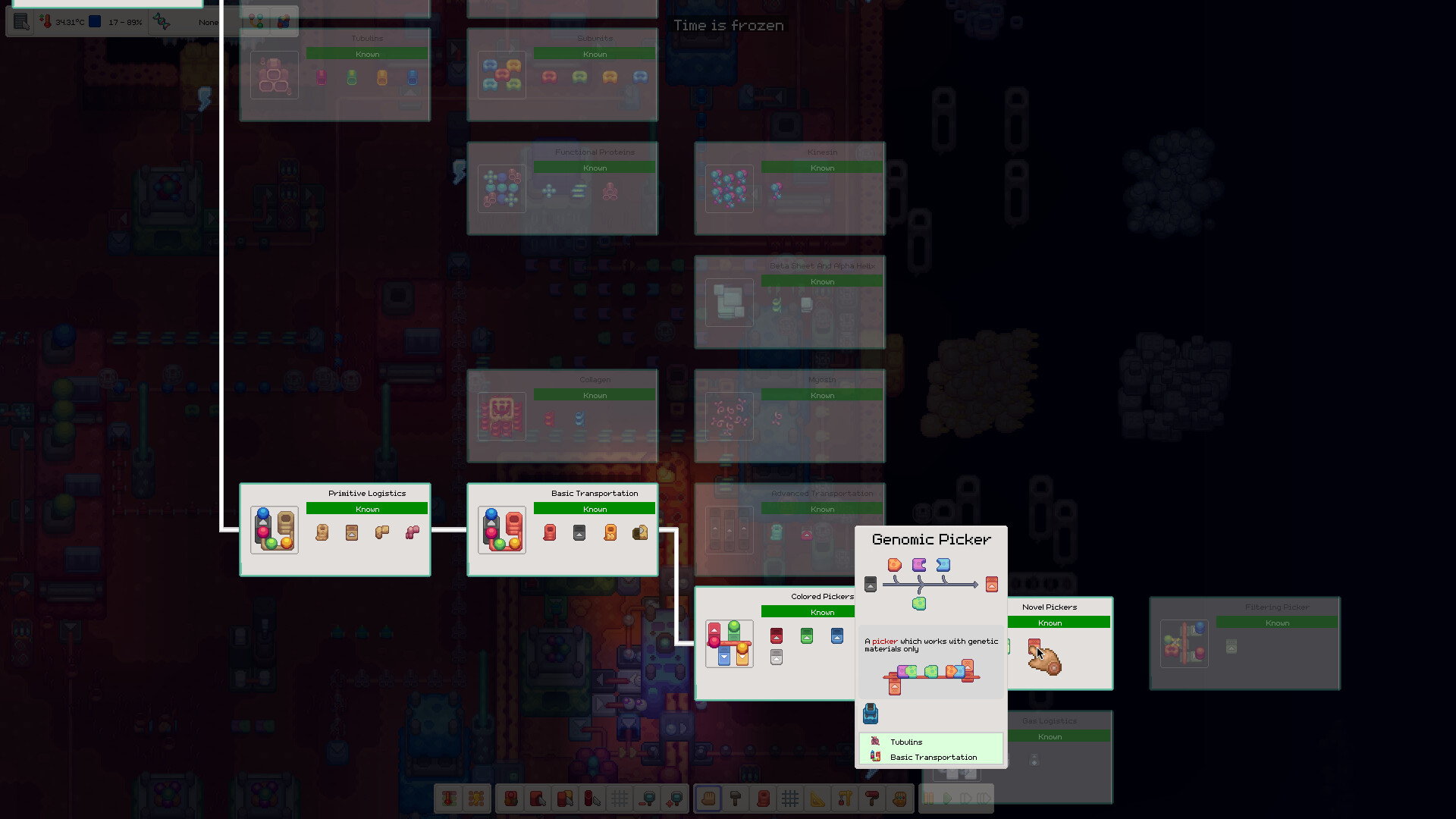

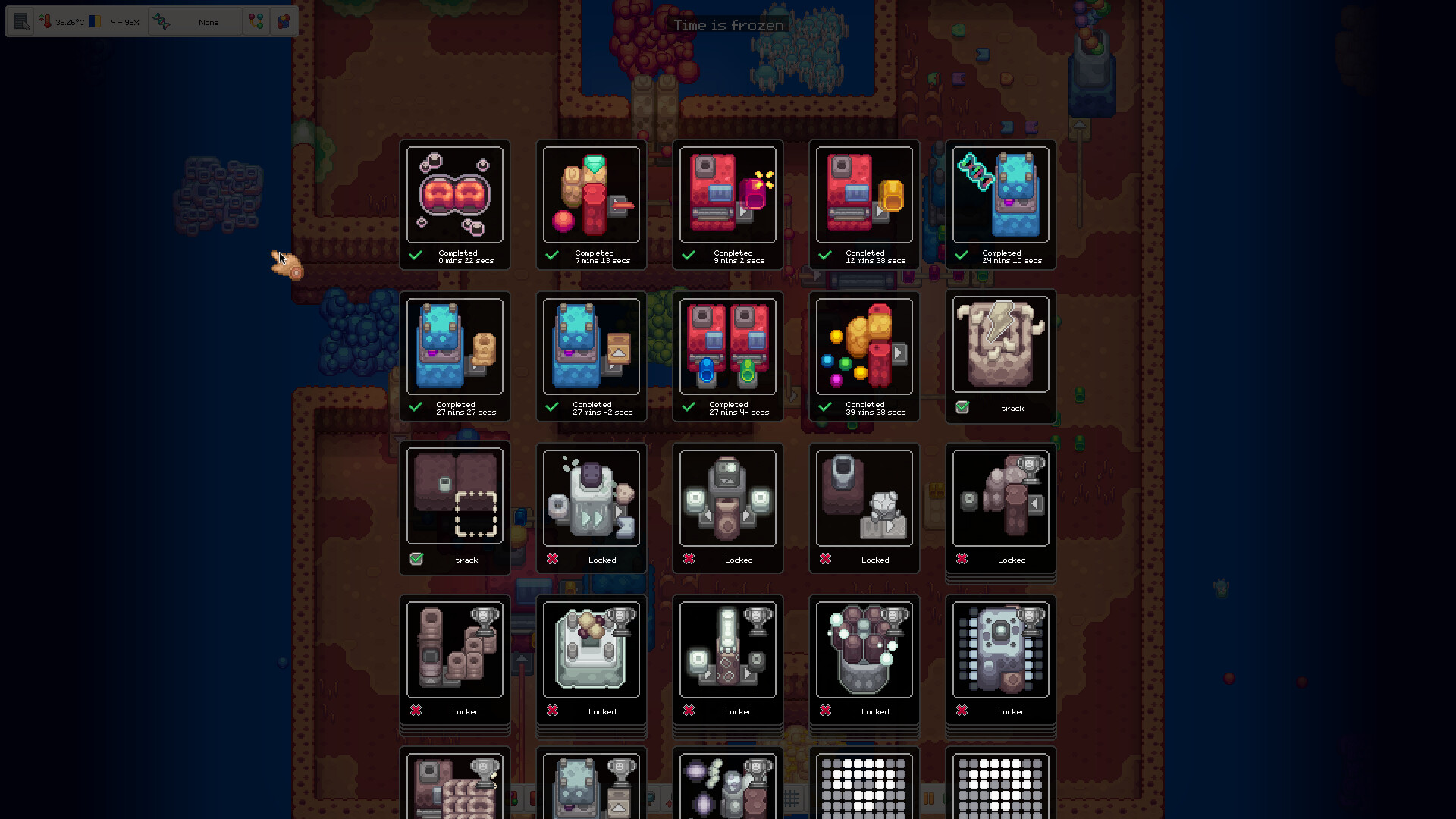


 Gallery
Gallery
 Changelog
Changelog
Development posts and patch notes from the official Lifecraft channels, following the work on circulation, digestion, blueprints, UI and modding.

Pathways documentation, extracellular circulation, game scale experiments and interactive tooltip work.
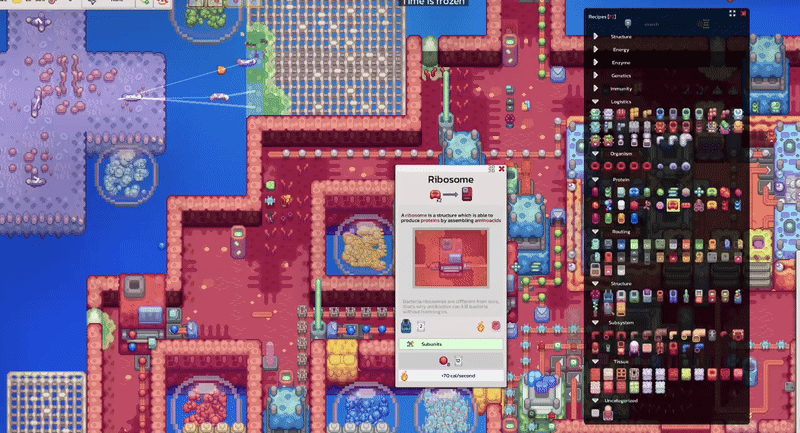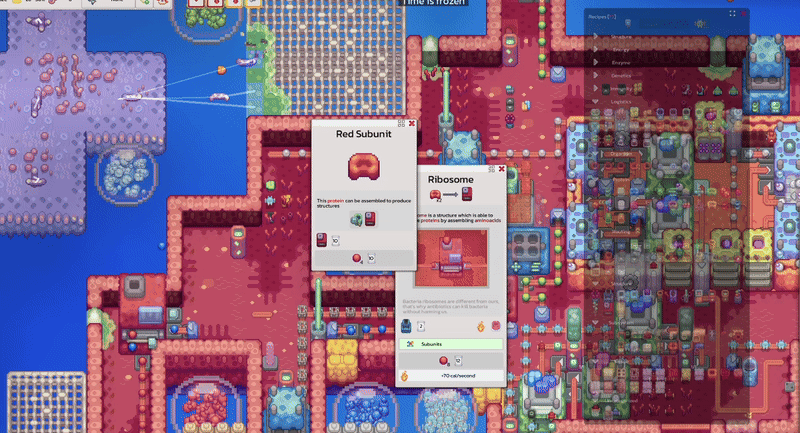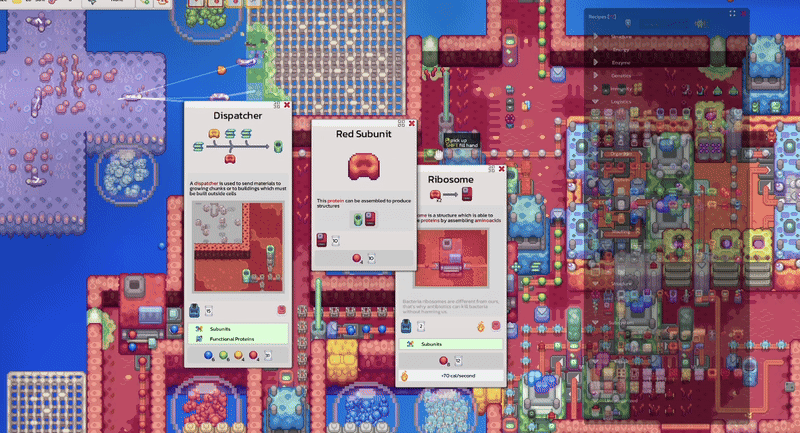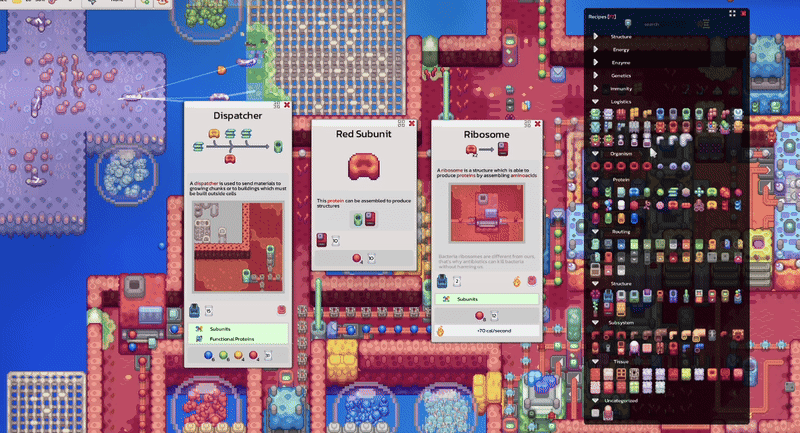
Pinnable tooltips, interface clarity improvements and new warm-up work around lipids.

A major update focused on blueprints, logic refactors, modding preparation and development fixes.
 Steam
Steam
Lifecraft is an Early Access project by Pixbits. The development posts invite players to share feedback while the organism keeps gaining new systems and mechanics.
 About Pixbits
About Pixbits
Pixbits is the duo behind Lifecraft and Junk Jack. Jack works on code and game design; XsX works on graphics, music and SFX.

